
Back Camp de forces (química) Catalan Kraftfeld (Computerphysik) German Campo de fuerza (química) Spanish میدانهای نیروی برهمکنشی (شیمی) Persian Champ de force (chimie) French Force field Italian 力場 (化学) Japanese Хүчний орон Mongolian Krachtveld (scheikunde) Dutch Pole siłowe (chemia) Polish
This article may require cleanup to meet Wikipedia's quality standards. The specific problem is: Grammar issues. (January 2024) |
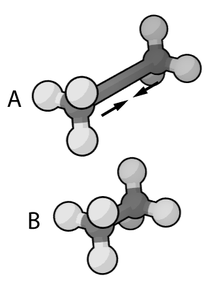
In the context of chemistry, molecular physics, physical chemistry, and molecular modelling, a force field is a computational model that is used to describe the forces between atoms (or collections of atoms) within molecules or between molecules as well as in crystals. Force fields are a variety of interatomic potentials. More precisely, the force field refers to the functional form and parameter sets used to calculate the potential energy of a system on the atomistic level. Force fields are usually used in molecular dynamics or Monte Carlo simulations. The parameters for a chosen energy function may be derived from classical laboratory experiment data, calculations in quantum mechanics, or both. Force fields utilize the same concept as force fields in classical physics, with the main difference being that the force field parameters in chemistry describe the energy landscape on the atomistic level. From a force field, the acting forces on every particle are derived as a gradient of the potential energy with respect to the particle coordinates.[1]
A large number of different force field types exist today (e.g. for organic molecules, ions, polymers, minerals, and metals). Depending on the material, different functional forms are usually chosen for the force fields since different types of atomistic interactions dominate the material behavior.
There are various criteria that can be used for categorizing force field parametrization strategies. An important differentiation is 'component-specific' and 'transferable'. For a component-specific parametrization, the considered force field is developed solely for describing a single given substance (e.g. water).[2] For a transferable force field, all or some parameters are designed as building blocks and become transferable/ applicable for different substances (e.g. methyl groups in alkane transferable force fields).[3] A different important differentiation addresses the physical structure of the models: All-atom force fields provide parameters for every type of atom in a system, including hydrogen, while united-atom interatomic potentials treat the hydrogen and carbon atoms in methyl groups and methylene bridges as one interaction center.[4][5] Coarse-grained potentials, which are often used in long-time simulations of macromolecules such as proteins, nucleic acids, and multi-component complexes, sacrifice chemical details for higher computing efficiency.[6]
- ^ Frenkel D (2007). Understanding molecular simulation : from algorithms to applications. Academic Press. ISBN 978-0-12-267351-1. OCLC 254835355.
- ^ Cite error: The named reference
:1was invoked but never defined (see the help page). - ^ Siu SW, Pluhackova K, Böckmann RA (April 2012). "Optimization of the OPLS-AA Force Field for Long Hydrocarbons". Journal of Chemical Theory and Computation. 8 (4): 1459–70. doi:10.1021/ct200908r. PMID 26596756.
- ^ Cite error: The named reference
Leach_2001was invoked but never defined (see the help page). - ^ Stephan, Simon; Horsch, Martin T.; Vrabec, Jadran; Hasse, Hans (2019-07-03). "MolMod – an open access database of force fields for molecular simulations of fluids". Molecular Simulation. 45 (10): 806–814. arXiv:1904.05206. doi:10.1080/08927022.2019.1601191. ISSN 0892-7022. S2CID 119199372.
- ^ Cite error: The named reference
Marrink_2007was invoked but never defined (see the help page).